
The detection and study of possible life forms beyond Earth must be
based on effectively minimal agnostic assumptions about the nature of the
organism. For example, the biochemistry, internal structure,
and composition of microbial life might be significantly
different from terrestrial microorganisms and cannot be predetermined. However, for life as we know it (and maybe as we don't know it!), three properties that we can reasonably assume as neccessary are: (A) an ability to reproduce itself in a self-regulated form, (B) the ability to maintain some degree of isolation of its internal compartment from the surrounding environment, and (C) it will require water and carbon.
Even though we are making some minimal assumptions, they are based on what is currently understood about the composition and chemistry of our solar sysem and probably the rest of the universe. Thus, MiDA is esentially an agnostic life detection instrument.
During
the process of reproduction, the organism's metabolism, mediated
by its membrane processes, will, by necessity change the surrounding
chemical and redox environment. Thus, to definitevly identify "life"
we must be able to detect such changes, rapidly and free
of extraneous or non-biogenic interferences. To help in this problem,
with support from NASA's Astrobiology Science & Technology
program (ASTID), we have developed a new phisico-chemically based detection instrument,
dubbed the Microbial Detection Array (MiDA). At the heart of MiDA is is an integrated electrochemically-based sensor array based on the Wet Chemistry Lab (WCL) sensor array onboard the successful 2007 Phoenix Lander mission [1].
|
Our proposed detection methodology is unique by making effectively agnostic "minimal assumptions" about life and by being designed to remove the effects of any chemical or physical interferences that might be mistaken for signs of life.
It is in-effect an agnostic life detection system. |
The MiDA is designed to provide a response to very small chemical and physical
changes occurring in one of two identical chambers via differentially
monitored integrated electrochemical sensor arrays consisting of ion selective electrodes (ISE) for inorganic ions, pH, oxydation reduction potential (ORP), and impedance spectroscopy/contactless electrical conductivity. Minimal metabolism will
alter the physico-chemical steady state in one chamber such that
a difference between the sensor arrays will result in a signal.
This detection system makes minimal assumptions about the nature
of the microorganism, assuming only that after addition of water
the it will reproduce and in the process cause changes in its
immediate surroundings by consuming or generating, metabolizing,
and transporting in both directions, a number of molecules and
ionic species.
A functional flow diagram of MiDA is shown below in Figure 1. The detection begins by placing a homogenized split-sample of
soil into each chamber, adding pure water, sterilizing at a high
temperature incompatible with any form of terrestrial life, and
zeroing. In the absence of any metabolically active organism in
either chamber, no signal will be detected. The "inoculation"
of one chamber with a minimal number of microorganisms, which
proceed to multiply, will produce a significant metabolically
generated disequilibrium in the system (compared to the control)
to provide a detectable signal. Replication of the experiment
and positive results would lead to the conclusion of biologically
induced changes. Changes resulting from non-biological chemical
reactions of whatever type are canceled out by the control. The
replication of the procedure, split sample, and minimal inoculation
protocol, eliminate non-biogenic causation. In addition to detecting
microbes, the sensor array will also characterize the chemical
composition and electrochemistry of the sample. MiDA is a differential
electrochemical sensor array that will provide the ability to
simultaneously monitor a specifically chosen set of chemical and
physical parameters in two identical “growth” chambers.
The sensitivity of the system provided by the two differentially
monitored sensor arrays (Figure 2-3) is such, that minimal
growth in one of the chambers will alter the chemistry and ionic
properties sufficiently to produce a difference between the two
sensor arrays and result in a measurable signal. This life detection
system makes minimal assumptions about the nature of any life
on Mars. It assumes only that, after addition of water, the microorganism
replicates and that in the process will produce small changes
in its immediate surroundings by consuming, metabolizing, and
excreting a number of molecules and/or ionic species [2-5].
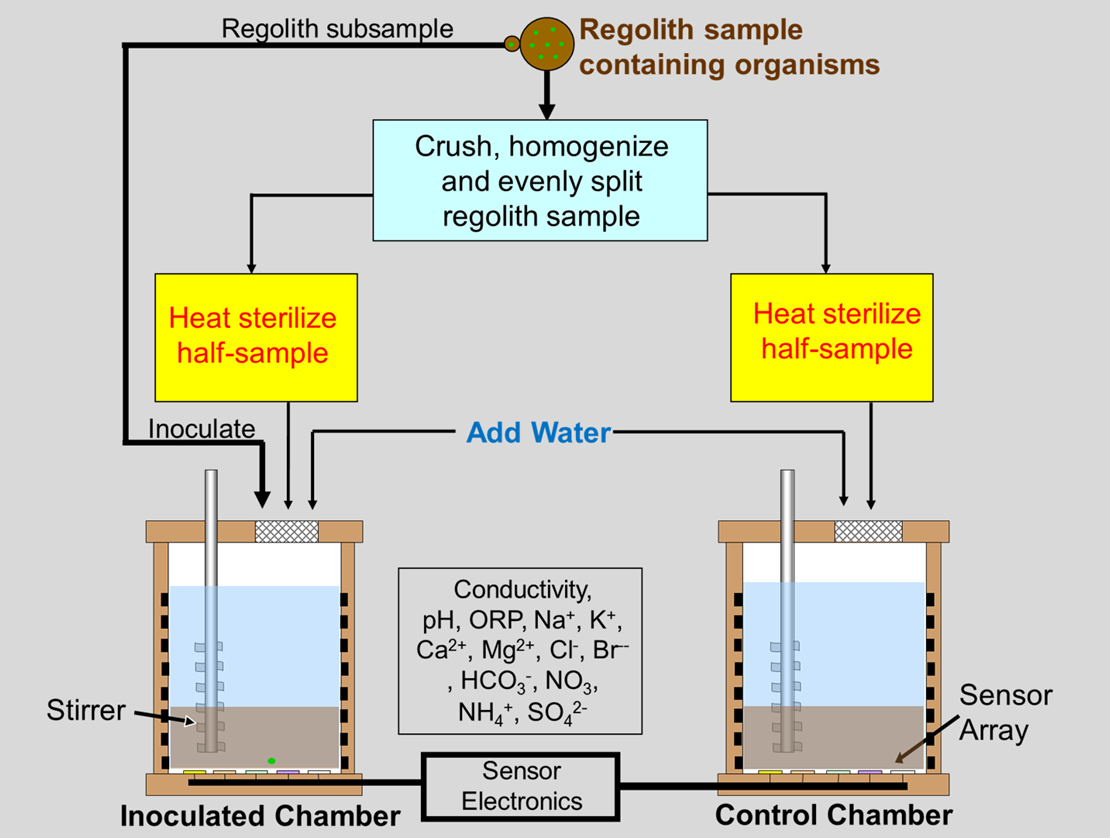
Figure 1. Functional Flow Diagram of MiDA.
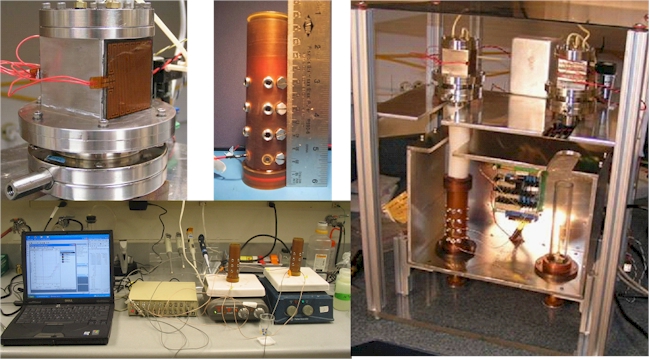

Figure 2. Prototype MiDA Test Unit
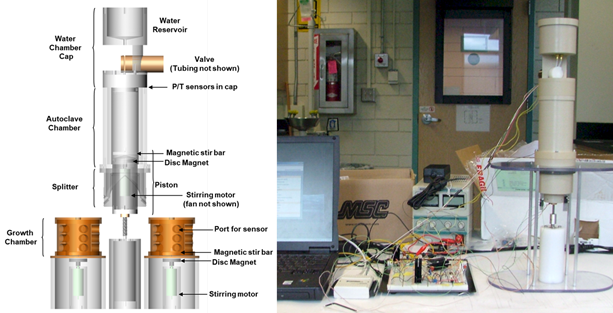
Figure 3. Mini-MiDA Fieldable Test Unit

|
[1] "The Wet Chemistry Experiments on the 2007 Phoenix Mars Scout Lander: Data Analysis and Results" S. P. Kounaves, et al., J. Geophys. Res., 2010, 115, E00E10, doi:10.1029/2008JE003084. Full Text
[2] "Microbial Life Detection With Minimal Assumptions", S. P. Kounaves, R.A. Noll, M.H. Hecht, M.G.Buehler, K. Lankford, S. West, in "Instruments, Methods and Missions for Astrobiology IV", R.B.Hoover, G.L.Levin, R.P. Paepe, A.Y.Rozanov (Eds), SPIE Proceedings, Vol. 4495, 137-144, 2002. Full Text PDF
[3] "Electrochemical Approaches for Chemical and Biological Analysis on Mars" S. P. Kounaves, ChemPhysChem, 2003, 4, 162-168. Full Text PDF
[4] "Microbial Detection Array (MDA) - Unambiguous Detection of Microbial Metabolic Activity in Astrobiology Applications", A. Hoehn, K. L. Lynch, J. Clawson, J. B. Freeman, J. Kapit, S. M. M. Young, S. P. Kounaves, and I. I. Brown, SAE Proceedings, ICES 2007, International Conference On Environmental Systems, Proceedings, Chicago, IL, USA, 2007SAE Document No. 2007-01-3190. Full Text PDF
[5] "A Method of Balancing Heat Sterilization with Minimal Media Degradation in Microbial Astrobiology Experiments" J. Kapit, MS Thesis, Tufts University, May 2009. Full Text PDF |
|